|

2009, Vol. 4 No. 1, Article 36
Differentation of closely related Vaccinal Strains of
Pasteurella multocida using Polymerase Chain Reaction(PCR)
Waheed ullah1, Muhammad Abubakar*2,
Muhammad Javed Arshed2,
Syed Muhammad Jamal2, Najma Ayub3 and Qurban
Ali2
1 Kohat University of Science and Technology, Kohat
2 National
Veterinary Laboratory, Park Road, Islamabad
3 Department of Microbiology, Quaid-e-Azam University, Islamabad
*Corresponding Author;
e-mail address: [email protected]
ABSTRACT
Nucleic acid based differentiation of closely related Pasteurella multocida vaccinal strains was performed. Morphological and biochemical characterization, HS-specific and species-specific PCR analysis of Pasteurella multocida vaccinal
strains were demonstrated useful in distinguishing hemorrhagic
septicemia-causing type B strains. The PCR assay performed for
species specific P. multocida by using primer pair KMT1T7 and
KMTISP6 resulted in amplification of all the strains. Another PCR
analysis carried out for H.S. causing strain conformation by using
primer pairs KTT72 and KTSP61 showed that only H.S. causing strains
were amplified. It was also observed that PCR amplification performed directly on bacterial colonies or cultures was an extremely rapid, sensitive method of P. multocida identification.
KEY
WORDS
Pasteurella multocida, Primers and PCR.
INTRODUCTION
Pasteurella multocida is a small, gram-negative, non–spore-forming cocco-bacillus with bipolar staining. It often exists as a commensal in the upper respiratory tract of many livestock, poultry, and domestic pet species and causes Haemorrhagic septicaemia (HS); a major disease of cattle and buffaloes occurring as catastrophic epizootics in many Asian and African countries, resulting in high mortality and morbidity. The disease has been recorded in wild mammals in several Asian and European countries. The Asian serotype B:2 and the African serotype E:2 (Carter and Heddleston system), corresponding to 6:B and 6:E (Namioka-carter system), are mainly responsible for the disease. In wild animals, serotype B:2,5 is predominantly present. The association of other serotypes, namely A:1, A:3 with a HS-like condition in cattle and buffaloes in India has been recorded (De-Alwis, 1990).
Vaccination against P. multocida can be achieved with whole-cell bacterins. However, efficacy is limited and restricted to the homologous serotype. Plain-bacterin, alum-type precipitated bacterin, and oil-adjuvant bacterin constitute the three widely used types of HS vaccine among which oil-adjuvant bacterin is the most effective.
A live intranasal vaccine prepared from a B:3,4 serotype of deer origin is also being used with reported success in southeast Asia (Muneer and Afzal, 1989).
Since the disease been reported even in the vaccinated animals and vaccination failure reports are common therefore the present study was undertaken with the objectives of; morphological and biochemical characterization of
P. multocida, HS-specific and species-specific PCR analysis of
Pasteurella multocida vaccinal strains to obtain a clear picture of HS vaccinal strains.
MATERIALS AND METHODS
Analysis of Pasteurella multocida
Three bovine vaccinal strains of P. multocida, B:2, B:6 and B:3,4 were investigated in this study. The vaccinal strains were taken from the vaccine seed bank of vaccine quality control section of National Veterinary Laboratory (NVL), Islamabad, Pakistan. The strains were inoculated intra-peritoneal into mice of 3-4weeks age. The mice were slaughtered within 6-7 hours and the mice heart and liver were cultured to obtain pure growth. Staining and biochemical tests (Catalase Test, Oxidase Test, Urease Production Test, H2S Production Test, Nitrate Reduction Test and Motility test) were performed in bacteriology laboratory of NVL to confirm
P. multocida before further investigation (Cheesbrough 1984).
Genomic DNA Analysis
Pure P. multocida culture was inoculated in brain heart infusion broth (BHI) at 37˚C for 24 hours. This culture was used for DNA extraction.
DNA Extraction Method (Antony et al., 2007)
The broth culture of P. multocida was transferred to an Eppendorff tube and centrifuged at 3000-x g for about 10 minutes. The pellet obtained after centrifugation was washed and re-suspended in PBS and then centrifuged again. The final pellet was re-suspended in 100 μl of distilled water. The mixture was boiled for 10 minutes in water bath and transferred immediately into ice and snap chilled for 30 minutes. The sample was then thawed and centrifuged at 3000 x g for 5 minutes. The supernatant was separated from pellet and used as template DNA.
PCR Analysis
1)
Agarose Gel Running:2% Agarose gel was prepared in Tris Boric EDTA (TBE) buffer. Ethidium bromide (0.1%) was added and was mixed gently. After polymerization of the gel, samples were added to fill the wells dip. The gel tank was filled with 1X TBE buffer up to the wells dip and run at a constant current of 80Volts. The final gel was viewed by UV trans-illumination.
2) PCR assay (Thermocycler, BioRed, PTA-200 with alpha engine) for analysis of both species-specific and HS-causing Type-B-specific
P. multocida was performed as follow:
a) Pasteurella multocida specific PCR Assay: The primer pair, KMT1SP6 and KMT1T7 was used which specifically amplified a product of approximately 460 base pair (bp) in all subspecies of
P. multocida (Townsend et al., 1998).
The primer sequences were:
KMT1SP6 5' –GCTGTAAACGAACTCGCCAC- 3'
KMT1T7 5' –ATCCGCTATTTACCCAGTGG- 3'
b) PCR Assay for HS-associated type B serotype of Pasteurella multocida:
PCR analysis for HS-associated type B serotype P. multocida identification was performed. Primer pair KTSP61 and KTT72 were used which specifically amplified a product of approximately 560 base pair (bp) in all HS causing serotype of
P. multocida.
The primer sequences were:
KTT72 5’-AGGCTCGTTTGGATTATGAAG-3'
KTSP61 5'-ATCCGCTAACACACTCTC-3'
The thermal cycling parameters included: The initial denaturation at 94˚C for 5 minutes; 30 cycles of 94˚C for 1 minutes, 53˚C for 1 minutes, and 72˚C for 1 minute; and final extension at 72˚C for 9 minutes.
RESULTS
Characterization of Pasteurella multocida
The study involved microbiological and molecular characterization of vaccinal strains namely B: 6, B: 2 and B: 3, 4. All the cultures showed luxuriant growth on blood agar (Oxoid) having translucent grayish or yellowish green colonies. All the microorganisms studied showed whitish gray rough opaque colonies on BHIA (Brain heart infusion Agar). The organisms when gram stained showed short rods that were gram-negative, coccobacilli, 0.2-0.4 mm in size. All the culture showing typical gram-negative coccobacilli were further processed through biochemical characterization (Table 1). The organisms when grown on TSA in the absence of CO2 at 37oC showed luxuriant growth. When inoculated in semisolid medium no motility was observed. These results were in close agreement with findings of Kumar et.
al. (2004).
DNA Extraction from Pasteurella multocida
The DNA extracted from all the three strains were run on agarose gel. All the samples gave positive bands for DNA presences. The first set of primer, KMT1T7 and KMT1SP6 was species specific and amplified all strains B:2, B:3,4 and B:6. of
Pasteurella multocida, corroborating the findings of Townsend et al., (1997).
In another reaction, PCR was carried for HS-causing strain conformation by using primer pair, KTT72 and KTSP61. In this reaction only HS-Causing strains were amplified while Non-HS-Causing was not amplified (Figures 1 & 2). The PCR assay was specific and sensitive. The concordance of PCR results with the defined toxigenic status indicates 100% specificity and sensitivity as described by Carol et al., (1998).
DISCUSSION
Discrimination of the B:2 serotype with the clone KMT1 requires additional hybridization analysis. However, present study revealed that oligonucleotide primers designed during nucleotide sequencing analysis of the clone 6b can be used to identify type B
P. multocida that causes HS (types B:2, B:5, and B:2,5). It is understood that this assay will not identify all HS-causing strains of
P. multocida, as these primers do not amplify DNA from type E:2 strains that cause HS in Africa. Townsend et al, (2001), the same results were obtained during present trial as the type B specific primers didn’t amplify the type B: 3, 4 (Figure 2).
The ability of the PCR assays described in this study to provide rapid identification of
P. multocida and confirmation of the HS-causing serotype has the potential to reform HS diagnosis in Southeast Asia. This technique could be implemented to rapidly confirm a field diagnosis of HS without the need to obtain pure cultures and perform extensive biochemical tests. The development of DNA based techniques for differentiation of serotypes could provide best facility as compared to conventional serotyping systems, and has a potential to overcome the deficiencies associated with current serotyping techniques, which depends on inconsistent expression of phenotypic traits.
REFERENCES
-
Antony P. X., G. K. Nair, V.
Jayaprakasan, M. Mini and T. V. Aravindakshan 2007. Nucleic acid
based differentiation of Pasteurella multocida serotypes. The
Internet Journal of Veterinary Medicine, 2 (2): 85-89.
-
Carol A, Lichtensteiger, Susan M,
Steenbergen, Ruby M, Lee, Dale D, Polson, Eric R. Vimr (1996).
Direct PCR analysis for toxigenic Pasteurella multocida. J clin.
Microbiol. 34:3035–3039.
-
Cheesbrough, M. 1984. Medical
laboratory manual for tropical countires. Vol. II, Microbiology.
ELBS, Butterworth, Cambridge.
-
De Alwis M.C.L. (1990):
Haemorrhagic Septicaemia. ACIAR Monograph No. 57, p. 36.
-
Hopkins B.A., Huang T.H.M. &
Olson L.D. (1998). Differentiating turkey post-vaccination
isolates of Pasteurella multocida using arbitrarily primed
polymerase chain reaction. Avian Dis., 42, 265-274.
-
Kumar A. A, Shivachandra S. B,
Biswas A, Singh V. P, Singh Vijendra P, Srivastava S. K (2004).
Prevalent serotypes of Pasteurella multocida isolated from
different animal and avian species in India. Vet. res. Commun
28:657-667
-
Muneer R, Afzal M (1989).
Preliminary studies on improved oil adjuvant vaccine for
haemorrhagic septicaemia. Revue Scientifique et Technique office
international des epizooties 8:999-1004.
-
Townsend KM, Dawkins HJ,
Papadimitriou JM (1997). REP-PCR analysis of Pasteurella
multocida isolates that cause hemorrhagic septicemia. Res Vet
Sci 63:151-5.
-
Townsend, K. M., A. J. Frost, C.
W. Lee, J. M. Papadimitriou and H. J. S. Dawkins, 1998.
Development of PCR Assays for Species and Type-Specific
Identification of Pasteurella multocida Isolates. J. Clinical
Microbiol., 36 (4): 1096–1100
-
Townsend, John D. Boyce, Jing Y.
Chung, Alan J. Frost, and Ben Adler (2001). Genetic organization
of Pasteurella multocida cap loci and development of multiplex
capsular PCR typing system. J. Clinical Microbiol., 39: 924-929.
Table 1
Microbiological and biochemical characterization of
Pasteurella multocida
|
Microbiological/biochemical properties
|
Reaction
|
|
Gram stain
|
-ve
|
|
Catalase
production
|
+ve
|
|
Haemolysis
|
-ve
|
|
Hydrogen
sulphide production
|
+ve
|
|
Urease
production
|
-ve
|
|
Oxidase production
|
+ve
|
|
Nitrate reduction
|
+ve
|
|
Indol reaction
|
+ve
|
|
Motility
|
-ve
|
|
Growth on MacConkey media
|
-ve
|
|
Growth in the absence of CO2
|
+ve
|
Fig. 1
PCR Results of amplification of both Non-HS-Causing and
HS-Causing vaccinal strains of
P. multocida but only HS-Causing strain is amplified
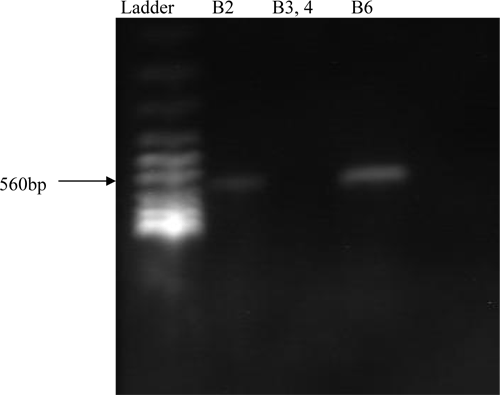
Fig. 2
Comparison of Band pattern of both Non-HS-Causing and
HS-Causing vaccinal strains
|
 |




